Moving (chemical) data around in a manner which allows its (automated) use in whichever context it finds itself must be a holy grail for all scientists and chemists. I posted earlier on the fragile nature of molecular diagrams making the journey between the editing program used to create them (say ChemDraw) and the Word processor used to place them into a context (say Microsoft office), via an intermediate storage area known as the clipboard. The round trip between the Macintosh (OS X) versions of these programs had been broken a little while, but it is now fixed! A small victory. This blog reports what happened when such a Mac-created Word document is sent to someone using Microsoft Windows as an OS (or vice versa).
Posts Tagged ‘opendata’
Data-round-tripping: moving chemical data around.
Saturday, November 20th, 2010For those of us who were around in 1985, an important chemical IT innovation occurred. We could acquire a computer which could be used to draw chemical structures in one application, and via a mysterious and mostly invisible entity called the clipboard, paste it into a word processor (it was called a Macintosh). Perchance even print the result on a laserprinter. Most students of the present age have no idea what we used to do before this innovation! Perhaps not in 1985, but at some stage shortly thereafter, and in effect without most people noticing, the return journey also started working, the so-called round trip. It seemed natural that a chemical structure diagram subjected to this treatment could still be chemically edited, and that it could make the round trip repeatedly. Little did we realise how fragile this round trip might be. Years later, the computer and its clipboard, the chemistry software, and the word processor had all moved on many generations (it is important to flag that three different vendors were involved, all using proprietary formats to weave their magic). And (on a Mac at least) the round-tripping no longer worked. Upon its return to (Chemdraw in this instance), it had been rendered inert, un-editable, and devoid of semantic meaning unless a human intervened. By the way, this process of data-loss is easily demonstrated even on this blog. The chemical diagrams you see here are similarly devoid of data, being merely bit-mapped JPG images. Which is why, on many of these posts, I put in the caption Click for 3D, which gives you access to the chemical data proper (in CML or other formats). And I throw in a digital repository identifier for good measure should you want a full dataset.
A Digital chemical repository – is it being used?
Tuesday, May 4th, 2010In this previous blog post I wrote about one way in which we have enhanced the journal article. Associated with that enhancement, and also sprinkled liberally throughout this blog, are links to a Digital Repository (if you want to read all about it, see DOI: 10.1021/ci7004737). It is a fairly specific repository for chemistry, with about 5000 entries. These are mostly the results of quantum mechanical calculations on molecules (together with a much smaller number of spectra, crystal structure and general document depositions). Today, with some help (thanks Matt!), I decided to take a look at how much use the repository was receiving.
On the importance of Digital repositories in Chemistry
Friday, April 3rd, 2009The preceeding blog entries contain stories about chemical behaviour. If you have clicked on the diagrams, you may even have gotten a Jmol view of the relevant molecules popping up. But if you are truly curious, you may even have the urge to acquire the relevant 3D information about the molecule, and play with it yourself. Even after 15 years of the (chemical) Web, this can be distressingly difficult to achieve (or can it be that it is only myself who wishes to view molecules in their native mode?). Thus the standard mechanism is to seek out on journal pages that disarming little entry entitled supporting information and to hope that you might find something useful embedded there. Embedded is the correct description, since the information is often found within the confines of an Acrobat file, and has to be extracted from there. Indeed, that is what I had to resort to in order to write one of the blog entries below. I ground my teeth whilst doing so.
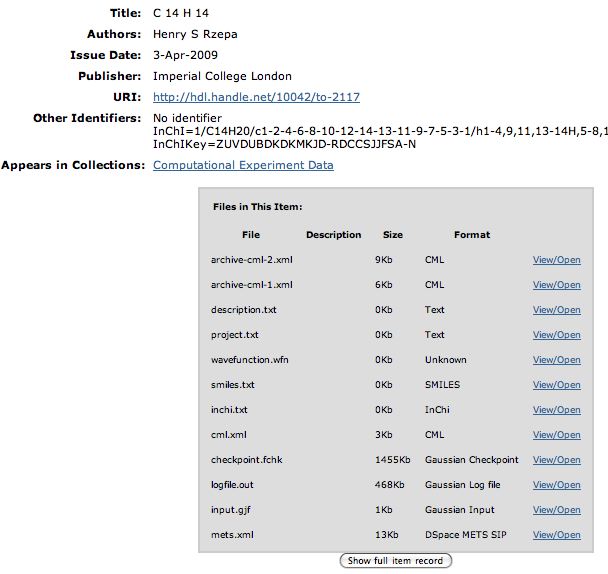
So is there a better way? We think so! The digital repository. If you click on this you should see the entry directly. What can you do there? Well, if you have suitable programs, you can download eg a Checkpoint file of the calculation that created the molecule model and re-activate it there. Or you can download just the CML file for viewing in any CML-compliant program (such as e.g. Jmol). Or you can check up on the InCHi string or the InChI Key of the molecule.