Posts Tagged ‘Acrobat’
Saturday, December 29th, 2018
The traditional structure of the research article has been honed and perfected for over 350 years by its custodians, the publishers of scientific journals. Nowadays, for some journals at least, it might be viewed as much as a profit centre as the perfected mechanism for scientific communication. Here I take a look at the components of such articles to try to envisage its future, with the focus on molecules and chemistry.
(more…)
Tags:Academic publishing, Acrobat, Articles, chemical discoveries, data, Data management, ELN, Information, Molecules, Narrative, PDF, Publishing, Research, Scholarly communication, Science, Scientific Journal, Scientific method, Technical communication, Technology/Internet, Web browser
Posted in Chemical IT | No Comments »
Monday, August 1st, 2016
In March, I posted from the ACS meeting in San Diego on the topic of Research data: Managing spectroscopy-NMR, and noted a talk by MestreLab Research on how a tool called Mpublish in the forthcoming release of their NMR analysis software Mestrenova could help. With that release now out, the opportunity arose to test the system.
(more…)
Tags:Acrobat, analysis software, chemical, Chemistry, City: San Diego, format type chemical/x-mnpub, media type, Mestrenova, non-commercial open software packages, Nuclear magnetic resonance, Nuclear magnetic resonance spectra database, Nuclear magnetic resonance spectroscopy, PDF, public key, Science, Scientific method, spectroscopy, Technology/Internet
Posted in Chemical IT | 3 Comments »
Saturday, July 19th, 2014
Whilst clusters of carbon atoms are well-known, my eye was caught by a recent article describing the detection of a cluster of boron atoms, B40 to be specific.[1] My interest was in how the σ and π-electrons were partitioned. In a C40, one can reliably predict that each carbon would contribute precisely one π-electron. But boron, being more electropositive, does not always play like that. Having one electron less per atom, one might imagine that a fullerene-like boron cluster would have no π-electrons. But the element has a propensity[2] to promote its σ-electrons into the π-manifold, leaving a σ-hole. So how many π-electrons does B40 have? These sorts of clusters are difficult to build using regular structure editors, and so coordinates are essential. The starting point for a set of coordinates with which to compute a wavefunction was the supporting information. Here is the relevant page: 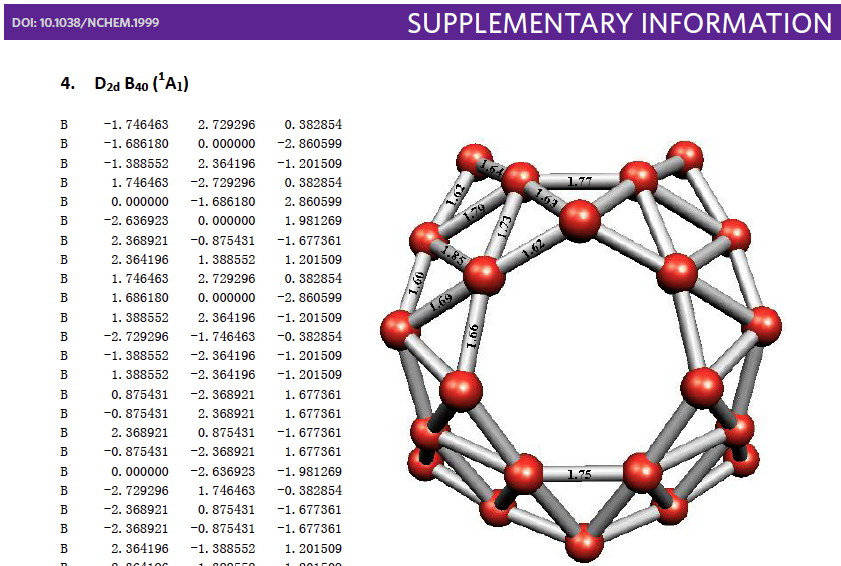 The coordinates are certainly there (that is not always the case), but you have to know a few tricks to make them usable.
The coordinates are certainly there (that is not always the case), but you have to know a few tricks to make them usable.
(more…)
References
- H. Zhai, Y. Zhao, W. Li, Q. Chen, H. Bai, H. Hu, Z.A. Piazza, W. Tian, H. Lu, Y. Wu, Y. Mu, G. Wei, Z. Liu, J. Li, S. Li, and L. Wang, "Observation of an all-boron fullerene", Nature Chemistry, vol. 6, pp. 727-731, 2014. https://doi.org/10.1038/nchem.1999
- H.S. Rzepa, "The distortivity of π-electrons in conjugated boron rings", Physical Chemistry Chemical Physics, vol. 11, pp. 10042, 2009. https://doi.org/10.1039/b911817a
Tags:Acrobat, Adobe, chemical markup, DOS, operating systems, PDF, pence, Unix
Posted in Chemical IT, Interesting chemistry | 2 Comments »
Monday, October 24th, 2011
I have for perhaps the last 25 years been urging publishers to recognise how science publishing could and should change. My latest thoughts are published in an article entitled “The past, present and future of Scientific discourse” (DOI: 10.1186/1758-2946-3-46). Here I take two articles, one published 58 years ago and one published last year, and attempt to reinvent some aspects. You can see the result for yourself (since this journal is laudably open access, and you will not need a subscription). The article is part of a special issue, arising from a one day symposium held in January 2011 entitled “Visions of a Semantic Molecular Future” in celebration of Peter Murray-Rust’s contributions over that period (go read all 15 articles on that theme in fact!).
(more…)
Tags:Acrobat, Amazon, Android, chemical structure drawing packages, e-books, HTML, HTML5, iPad, iPads, Java, KF8, Kindle, mobile devices, opendata, Peter Murray-Rust, printing costs, SVG, Vector Graphics
Posted in Chemical IT, General | 3 Comments »
Monday, September 7th, 2009
The science journal is generally acknowledged as first appearing around 1665 with the Philosophical Transactions of the Royal Society in London and (simultaneously) the French Academy of Sciences in Paris. By the turn of the millennium, around 10,000 science and medical journals were estimated to exist. By then, the Web had been around for a decade, and most journals had responded to this new medium by re-inventing themselves for it. For most part, they adopted a format which emulated paper (Acrobat), with a few embellishments (such as making the text fully searchable) and then used the Web to deliver this new reformulation of the journal. Otherwise, Robert Hooke would have easily recognized the medium he helped found in the 17th century.
(more…)
Tags:A. I. Magee, A. Jana, A. P. Dove, Acrobat, American Chemical Society, aqueous solution, Balasundaram Lavan, C. S. M Allan, C. Wentrup, Chemical IT, chemical plugin, Chemoinformatics, Colorado, D. A. Widdowson, D. C. Braddock, D. J. Williams, D. R. Carbery, D. Scheschkewitz, Dalton Trans, digital Acrobat, E. H. Smith, E. M. Barreiro, E. W. Tate, Enhance Chemical Electronic Publishing, Extrusion Reactions, F. Diederich, F. Santoro, French Academy, G. Siligardi, G. Stammler, Ge, H. S. Rzepa, HTML, I. Omlor, I. Pavlakos, Interchange Apical, Interesting chemistry, Ion-Pair Mechanisms, β-diketiminate metal alkoxides, J. Clarke, J. Jana, J. L. Arbour, J. Lorenzo Alonso-Gómez, J. P. White, J. R. Arendorf, journal editor, K. K. (Mimi) Hii, K. P. Tellmann, King, Kuok Hii, L. A. Adrio, L. Johannissen, Lewis Base Catalyst, M. E. Cass, M. Hii, M. J. Cowley, M. J. Fuchter, M. J. Harvey, M. J. Humphries, M. J. Porter, M. Jakt, M. R. Crittall, M. Ritzefeld, M. Weimar, Marshall, Michael Wright, N. Berova, N. Harada, N. J. Mason, N. Mason, N. Masumoto, O. Casher, opendata, P. G. Pringle, P. Jutzi, P. Lo, P. Seiler, Paris, Peter Murray-Rust, polymerization, Porter, printing, R. B. Moreno, R. M. Williams, R. Schleyer, R. Wilhelm, Rappaport, RDF, representative, Robert Hooke, Royal Society in London, S. Díez-González, S. Lai, S. M. Allan, S. Martin-Santamaria, Sonsoles Martên-Santamarêa, Square Pyramidal Molecules, T. Lanyon-Hogg, the Philosophical Transactions of the Royal Society, V, V(III) Co, V. C. Gibson, V. Huch, V. W. Pike, W. B. Motherwell, Web Application, Web Table, XML, XSLT, Ya-Pei Lo
Posted in Chemical IT, Interesting chemistry | 6 Comments »
Friday, April 3rd, 2009
The preceeding blog entries contain stories about chemical behaviour. If you have clicked on the diagrams, you may even have gotten a Jmol view of the relevant molecules popping up. But if you are truly curious, you may even have the urge to acquire the relevant 3D information about the molecule, and play with it yourself. Even after 15 years of the (chemical) Web, this can be distressingly difficult to achieve (or can it be that it is only myself who wishes to view molecules in their native mode?). Thus the standard mechanism is to seek out on journal pages that disarming little entry entitled supporting information and to hope that you might find something useful embedded there. Embedded is the correct description, since the information is often found within the confines of an Acrobat file, and has to be extracted from there. Indeed, that is what I had to resort to in order to write one of the blog entries below. I ground my teeth whilst doing so.
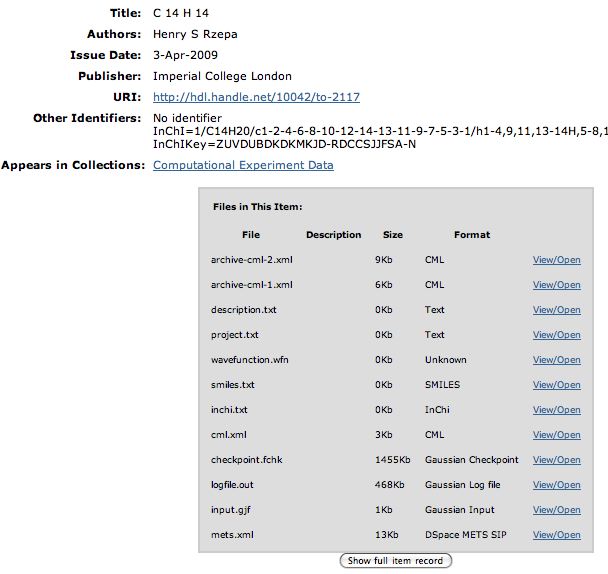
So is there a better way? We think so! The digital repository. If you click on this you should see the entry directly. What can you do there? Well, if you have suitable programs, you can download eg a Checkpoint file of the calculation that created the molecule model and re-activate it there. Or you can download just the CML file for viewing in any CML-compliant program (such as e.g. Jmol). Or you can check up on the InCHi string or the InChI Key of the molecule.
(more…)
Tags:Acrobat, Checkpoint, chemical, chemical behaviour, Chemical IT, editor in chief, opendata, Peter Murray-Rust, Web-enhanced tables
Posted in Chemical IT | No Comments »

