Most of the chemical structure diagrams in this blog originate from Chemdraw, which seems to have been around since the dawn of personal computers! I have tended to use this program to produce JPG bitmaps for the blog, writing them out in 4x magnification, so that they can be scaled down for display whilst retaining some measure of higher resolution if needed for other purposes. These other purposes might be for e.g. the production of e-books (using Calibre), the interesting Blog(e)book format offered as a service by Feedfabrik, or display on mobile tablets where the touch-zoom metaphor to magnify works particularly well. But bitmap images are not really well future proofed for such new uses. Here I explore one solution to this issue.
I have previously mentioned scalable vector graphics (SVG) as an alternative, and fortunately the production of such has become routine.3 The diagram above2 is indeed SVG (and if you cannot see it, then try a modern SVG-capable browser1). It was produced thus:
- Drawn in Chemdraw
- Exported as Encapsulated postscript
- Imported into Scribus, an Open Source desktop publishing program (where it can be annotated/edited if need be)
- This program will also need Ghostscript installed to handle the EPS
- and exported from Scribus to SVG.
- Notice how the diagram above automatically scales to fill the width of the page. If you click on it, you get the diagram on its own. If you zoom the browser window, it should scale perfectly.
- I note that these SVG diagrams work well in e-books or blogbooks.
1 I notice that Internet Explorer 9 (both 32- and 64-bit versions) will display (and save) the above diagram if you click on it, but it cannot (yet) be inlined into the post, although the documentation implies it should.
2 The version below is the conventional JPG form (click on it to see the original 4x version).
3. Historical note. Peter Murray-Rust and I have been promoting SVG for use in chemistry for 11+ years now. For one ancient page, see here. The syntax has decayed somewhat, but some of the diagrams still work!
Tags: chemical diagrams, chemical structure diagrams, desktop publishing, e-books, Feedfabrik, GIF, GPS, Internet Explorer, Peter Murray-Rust, software environment, Vector Graphics
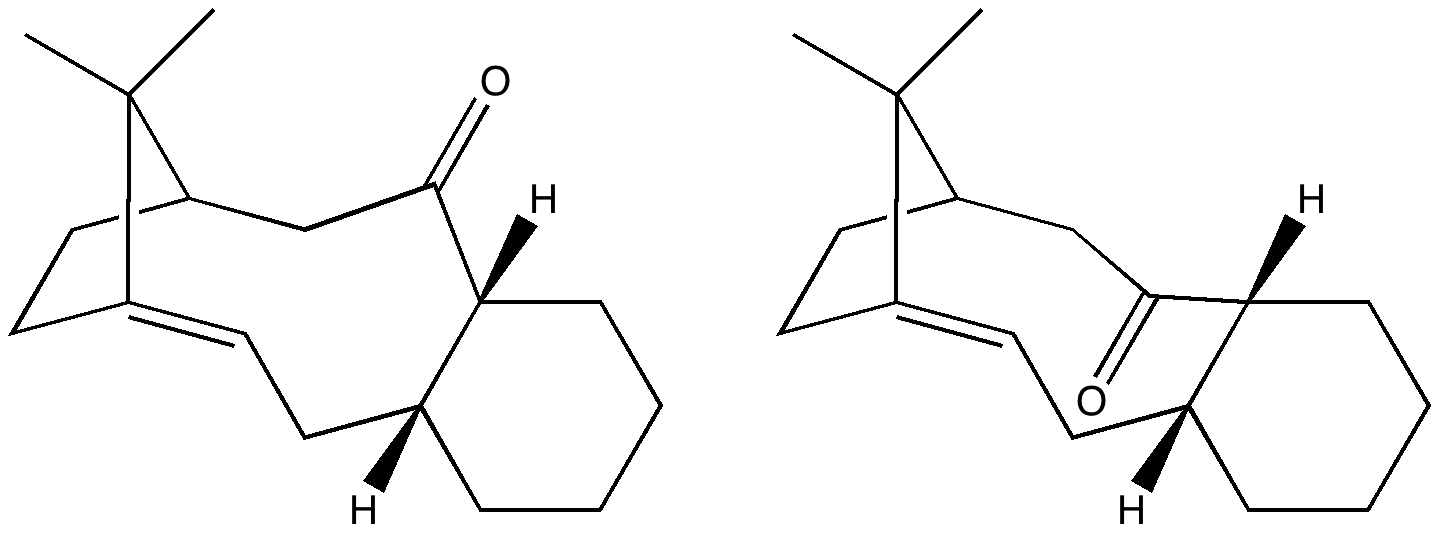
Hrm, maybe you could’ve published the original ChemDraw JPG for side-by-side examination. The carbonyl group looks kinda odd to me (too long double bond, to small oxygen “O”), but maybe I am just making it up.
As per request, see above. No re-touching! I am reminded of the story about Napier’s logarithms, whereby he inserted small deliberate (and inconsequential) errors into his log tables to catch out people who might have been inclined to re-publish them. There are rumours that chemical structure drawing programs also have tiny features (mountweazels) which were used to detect anyone who was publishing using the program, but without a valid license. But these are just stories, and probably untrue.
Hi Henry,
You may already know this, but ChemDoodle can output directly to SVG and several other graphical standards. In addition, ChemDoodle can create HTML5 ChemDoodle Web Components for dynamic chemical content; see Interactive chemistry ebooks: interactive figures
Many publishers and formats are now embracing HTML5, such as Amazon (Amazon introduces new HTML5-based eBook format), and we should start seeing WebGL on the iPad and other mobile devices soon, so it is definitely worth the effort to invest in these new technologies.
In the next update, ChemDoodle SVG output will contain chemical metadata, so that round-tripping that information will be possible. Even if other providers will not be supporting technologies like SVG, we certainly will be.
Greetings Kevin.
Yes, regarding Amazon, I did blog this and also ChemDoodle.
As for the round tripping, do please use CML. As I see it, you currently express data for ChemDoodle via line escaped Molfiles (the need to insert \n at the end of each line of a mol file only illustrates how old that format actually is!)
By the way, there is a very impressive demo of Jmol on an Android tablet, produced by Bob Hanson. I do not have such a beast nearby. Does anyone know if it supports WebGL (and indeed SVG)?
Actually, the ChemDoodle Web Components library reads many formats, including CML. We use MOL files for the default instructions because they are typically smaller in size. The use of the \n would be necessary for CML input as well, because the Javascript string cannot contain new lines. Of course, we are also open to new suggestions for handling CML.
ChemDoodle desktop reads and writes CML (v1 and v2).
I think the important note though, is that if you want to see these technologies adopted and used, then you have to support software that supports them, especially if the software you have used for decades will not.
Actually, removing the line breaks from a CML file should NOT break it (unlike a Molfile). Shall we try the experiment?
The above is all on one line!
Regarding adoption of new technologies, by far the greatest use of programs such as Chemdraw here is for including high quality printable diagrams into Microsoft Word and Powerpoint. Whilst M$’s IE9 browser might support eg HTML5/SVG, I think Office 2011/2012 probably does not.
Do enlighten me Kevin on how Chemdoodle might be used in such an environment? If it can, lots of people here might be convincible (but in fact most chemists are ultra conservative when it comes to this sort of thing).
First, Henry, I have followed your work since I first became interested in computers in chemistry, and have always been and always will be a huge fan. I take your standards seriously, and we follow them in all of our products (from CML to MIME types).
You are correct about XML not requiring return lines. I was under the false impression that processing instructions (such as the declaration) must be followed by return lines. So definitely, CML is a very good choice for those that do not want to deal with such syntax issues with the ChemDoodle Web Components.
About Microsoft Word and Powerpoint, the ChemDoodle Web Components would not be such a great tool, but ChemDoodle desktop certainly is. The ChemDoodle Web Components is a Javascript library for creating HTML5 content for chemistry and other sciences. ChemDoodle desktop is an end user software application for creating chemical graphics and other chemical publishing functions. ChemDoodle is compatible with Microsoft Office, Apple iWork and OpenOffice (and other variants), as well as other third party products like Adobe software. You can paste scalable vector graphics directly into these software suites on Windows, Mac OS X and Linux. On Mac OS X, we also support round-trip editing with most of these applications, including Microsoft Office 2011 and iWork Keynote. We will also be supporting round-trip editing on Windows soon via OLE.
If you do not mind me stealing a bit of your time, I would be very happy to speak with you some more about your work and the future of this technology.
My previous example of ChemDoodle used a Molfile with the linefeeds replaced by \n, and with the hard wraps removed to enable it to be specified in a Javascript array. We generate CML from a High-performance computing facility linked to a DSpace digital repository, and here the conversion to CML is carried out automatically. It would be trivial to write this file without hard wraps, and thence to use it directly for e.g. ChemDoodle. Would the current version support this directly? One can also insert CML into SVG, and have only the CML namespace read by Chemdoodle (although the array would probably now contain e.g. more than 10,000 characters). It would be fantastic Kevin if you could come up with a demo, which could be routinely used on e.g. this blog.
The other operation which I think is needed is the ability to extract e.g. the CML from the ChemDoodle sandbox and re-use it elsewhere. One aspect of e.g. the IOS model on iPad is that the data is sandboxed into the app, and it can be quite cumbersome to extract it from there and inject a copy into another sandbox. I gather Android is less restrictive (and perhaps also less secure) in this aspect?
So is demo of ChemDoodle possible which uses a data array containing CML, and some mechanism for extracting just that data out of the array and depositing it onto a clipboard (or even a file) or other sandbox? Is that breaking all the rules?
Sure, let me put together a demo with CML and the ChemDoodle Web Components. I will put it up when I get a chance tonight.
There are two ways to deal with CML, since it is XML. The first is by Javascript string, and we would then need a Javascript XML parser. To get around this, we use a webservice to parse the CML as will be shown in the demo. XHR2 is used so that anyone can call this directly from any website. The second way is via the HTML DOM, if you wanted to embed the CML in an SVG element or somewhere else in the DOM. Currently, the ChemDoodle Web Components doesn’t do this, but maybe in the future.
Any clipboard functions, including copy and paste, are forbidden in Javascript, so a simple way to extract the data is not possible. However, using the XHR2 service, we can save the data to a file format of your choice, and then have the user download it. This should also work on Android, but probably not iOS. I will show this in the demo as well.
Here is the demo: http://web.chemdoodle.com/demos/chemical-markup-language-cml
[…] Henry Rzepa Chemistry with a twist « Blogbooks, e-books and future proofing chemical diagrams. […]