Scientific data in chemistry has come a long way in the last few decades. Originally entangled into scientific articles in the form of tables of numbers or diagrams, it was (partially) disentangled into supporting information when journals became electronic in the late 1990s.[1] The next phase was the introduction of data repositories in the early naughties. Now associated with innovative commercial companies such as Figshare and later the non-commercial Zenodo, such repositories have also spread to institutional form such as eg the earlier SPECTRa project of 2006[2] and still evolving.[3] Perhaps the best known, and certainly one of the oldest examples of curated structural data in chemistry is the CCDC (Cambridge crystallographic data centre) CSD (Cambridge structural database) which has been operating for more than 55 years now, even before the online era! Curation here is the important context, since there you will find crystal diffraction data which has been refined into a structural model, firstly by the authors reporting the structure and then by CSD who amongst other operations, validate the associated data using a utility called CheckCIF.[4] What perhaps is not realised by most users of this data source is that the original or “raw” data, as obtained from a X-ray diffractometer and which the CSD data is derived from, is not actually available from the CSD. This primary form of crystallographic data is the topic of this post.
Most chemical data now emerges from an instrument, where it is already partially processed internally before being offered. Such raw/primary data is perhaps best known in the form of NMR information, where it is offered by the instrument in the form of an FID or free induction decay. Its transformation from this form into what all chemists know as a spectrum requires further software processing, and including other operations such as peak integration. It is this processed spectrum that had traditionally been offered as part of a scientific article (often only in visual, or peak listed form) and rarely has the FID form been made available to anyone interested. It is important to state that the transformation to spectrum also incurrs significant loss of data. An interesting project led by the editors of two organic chemistry journals[5],[6] had the aim of encouraging the submission of FAIR data to the journal, although in fact the project actually concentrated on the submission of raw NMR data. As it turned out, only a very small proportion of all the submissions to these journals over the period of a year actually provided such data (~113 datasets) in the form of ZIP archives‡ and containing anywhere between one and ~100 actual sets of raw NMR data per archive. One should make the point that raw data is not necessarily FAIR data. The latter requires rich metadata describing the data to become findable, accessible, interoperable and reusable (FAIR), and such metadata was not actually generated as part of this publisher project.
Here I will take a closer look at potentially FAIR raw data in the area of crystallography. This project is perhaps less well known than the previous one,[5],[6] hence the present post strives to make it better known. As with NMR, a useful starting point is to describe the various stages in the lifecycle of crystal data.
- A crystal is mounted in the diffractometer and x-ray diffraction images are recorded. These are considered the raw data, and as with most instruments, their form is determined both by the instrument itself and the software used to start the refinement process into a molecular structure.
- This refinement then assigns a space group to the data and derives so-called structure factors or hkl data. This data can now be captured in a much more standard form known as a CIF (crystallographic information file) and is nowadays the format that is deposited with CSD.
- A reduced form of the CIF file, containing a sub-set of the information but lacking the hkl data is much the more common, and was the form originally sent to CSD until a few years ago.
- Very often an image of the resulting model for the molecular structure is also included. Whilst it is based on the data in the CIF file, it does not contain reusable data as such and is considered as being made available only for human use and perception.
It is form 1 that is missing from the CSD datasets. Because it can be quite large (~0.5-9 Gbyte), the current recommendation is that it is not stored on the CSD but on local data repositories.† So now we see a need to establish if possible bidirectional links between type 1 and types 2-4 and to identify what characteristics of FAIR each has. Primarily, the F (findable) of FAIR will be explored here. This is done by illustrating some searches for this data, based on the metadata registered for it with DataCite.
- https://commons.datacite.org/?query=relatedIdentifiers.relatedIdentifier:10.5517* (157 works)
This simple search identifies any entry in any repository which cites in its metadata record the DOI for an entry in CSD, taking the form 10.5517* which is common to all entries. - ?query=relatedIdentifiers.relatedIdentifier:*10.5517*+AND+(media.media_type:chemical/x-cif+OR+media.media_type:application/x-7z-compressed+OR+media.media_type:application/gzip+OR+media.media_type:application/zip) (9 works).
This also specifies that search 5 is further constrained by requiring one of four media types to ALSO be present in the repository metadata record. These types are standard compressed archives which the raw crystal data is likely to be stored as, along with a CIF entry that is clearly associated with crystal structure data. The Boolean OR indicates that any one of them can be present! One can now be a little more certain that these entries contain crystal structure data. That we cannot be absolutely certain is clearly a current deficiency of the metadata present for the entries! - ?query=identifier:*10.5517*+AND+(relatedIdentifiers.relatedIdentifier:*10.14469*) (7 works)
Eight works from search 6 originate from a repository with the prefix 10.14469* and so now one can reverse the direction and ask how many are referenced in the metadata for each published item in the CSD? Around 945,473 entries in the CSD currently have a persistent DOI identifier associated with them, all starting with 10.5517* and so now one can search for how many of these also reference a related identifier at 10.14469* Seven of them show up there. - Also in the CSD metadata records is an item with the attribute relationType=”IsDerivedFrom” carrying the meaning that the CSD data is itself derived from (raw) data held elsewhere. This information is captured during the deposition process with CCDC as per below.
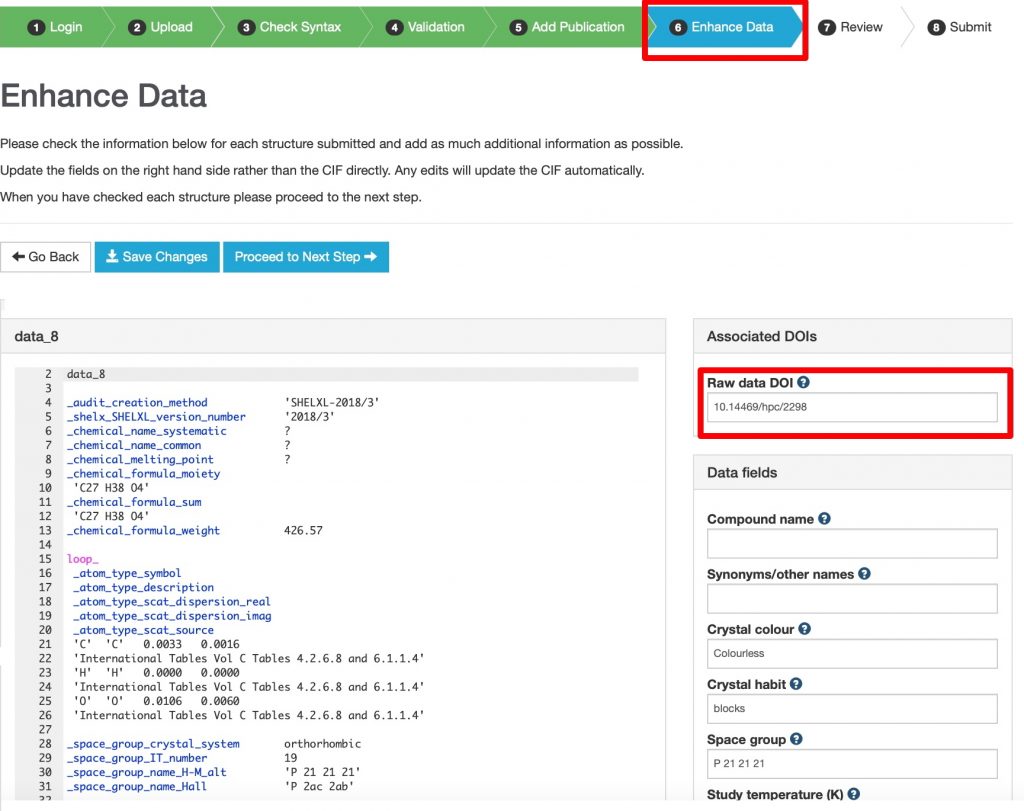
https://commons.datacite.org/?query=identifier:*10.5517*+AND+(relatedIdentifiers.relationType:IsSourceOf+OR+relatedIdentifiers.relationType:IsDerivedFrom) (7 works)
This constrains to datasets at CSD that are associated with additional raw data by IsDerivedFrom or IsSourceOf relationships.♥ CCDC tell me the true number is around 65 so the origins of this mismatch need to be identified.
So projects aiming to capture data from chemical instrumentation are just starting to reveal the potential of this modern system for storing data in two or more locations and reconciling various forms of this data, from raw form to derived or processed data. The interested user can then use whichever form is most relevant to their needs, and having found one form can then trace back to the other form(s). We might anticipate many developments in this area in the near future.
‡One has to expand the archive to find out how many actual raw datasets are inside, rather than knowing beforehand how many datasets are contained there, or anything else about their properties. †The publication process is described here for one repository at DOI: 10.14469/hpc/10178 ♥From the DataCite schema; <relatedIdentifier relationType="IsDerivedFrom">... </relatedIdentifier> IsDerivedFrom should be used for a resource that is a derivative of an original resource. In this example, the dataset is derived from a larger dataset and data values have been manipulated from their original state. <relatedIdentifier relationType="IsSourceOf">... </relatedIdentifier> IsSourceOf is the original resource from which a derivative resource was created. In this example, this is the original dataset without value manipulation.
This post has DOI: 10.14469/hpc/10177
References
- A.M. Hunter, and A.B. Smith, "Review of Supporting Information at <i>Organic Letters</i>", Organic Letters, vol. 17, pp. 2867-2869, 2015. https://doi.org/10.1021/acs.orglett.5b01700
- J. Downing, P. Murray-Rust, A.P. Tonge, P. Morgan, H.S. Rzepa, F. Cotterill, N. Day, and M.J. Harvey, "SPECTRa: The Deposition and Validation of Primary Chemistry Research Data in Digital Repositories", Journal of Chemical Information and Modeling, vol. 48, pp. 1571-1581, 2008. https://doi.org/10.1021/ci7004737
- M.J. Harvey, A. McLean, and H.S. Rzepa, "A metadata-driven approach to data repository design", Journal of Cheminformatics, vol. 9, 2017. https://doi.org/10.1186/s13321-017-0190-6
- A.L. Spek, "Structure validation in chemical crystallography", Acta Crystallographica Section D Biological Crystallography, vol. 65, pp. 148-155, 2009. https://doi.org/10.1107/s090744490804362x
- A.M. Hunter, E.M. Carreira, and S.J. Miller, "Encouraging Submission of FAIR Data at <i>The Journal of Organic Chemistry</i> and <i>Organic Letters</i>", The Journal of Organic Chemistry, vol. 85, pp. 1773-1774, 2020. https://doi.org/10.1021/acs.joc.0c00248
- A.M. Hunter, E.M. Carreira, and S.J. Miller, "Encouraging Submission of FAIR Data at <i>The Journal of Organic Chemistry</i> and <i>Organic Letters</i>", Organic Letters, vol. 22, pp. 1231-1232, 2020. https://doi.org/10.1021/acs.orglett.0c00383
This post dates from March 1st, at which time the search https://commons.datacite.org/?query=identifier:*10.5517*+AND+(relatedIdentifiers.relationType:IsSourceOf+OR+relatedIdentifiers.relationType:IsDerivedFrom) returned 7 hits, whilst around 65 were expected from internal records of the CCDC. A little while later the results reduced to 0, as a result of temporary internal changes in the metadata registration at CCDC. Now I can report that with the completion of this refactoring, the same search returns 216 hits as of today.
Deposition of raw image data for crystal structures seems to have been driven until now by authors, and so the 216 hits are probably a natural and uncoordinated emergence of the provision of such data. One might imagine that when journals, their referees, CCDC itself and agencies such as IUPAC start to promote and encourage the deposition of primary research data, this number will swell considerably. It might be interesting to repeat this search at periodic intervals to see if this is actually happening.