For obvious reasons, anti-viral molecules are very much in the news at the moment. Thus Derek Lowe highlights Molnupiravir which is shown as a hydroxylamine, the representation originating from the Wikipedia page on the molecule.
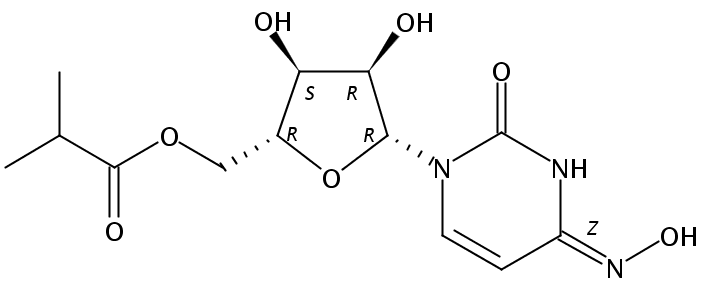
I like stereocentres more clearly identified using eg R/S notation and so I went to another source of information, SciFinder, which represents the molecule in a different way. There you get the stereocentres unambiguously identified for you, but the hydroxylamine is now replaced by an oxime!
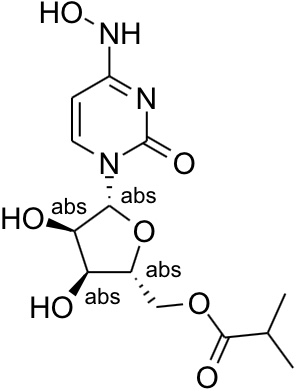
The Reaxys database renders it differently again as below (note the different rotamer for the hydroxylamine):
Are they all talking about the same molecule? Well yes, since the hydroxylamine and the oxime are related by tautomerism, but it takes a bit of effort to fully reconcile these three representations with each other. So the next question is does it matter which tautomer is selected to represent a molecule? Since they differ by proton transfers between acidic atoms (N,O) the presumption is that the equilibrium between the tautomers is fast and so the predominant species is determined by the position of the equilibrium. The tautomer matters in another sense. This molecule clearly interacts with DNA, and very probably by paired hydrogen bonding. It is this hydrogen bonding that was crucial in helping Watson and Crick to postulate the first successful model of DNA itself,[1] famously enabled by them using the correct tautomeric form of the component DNA bases! So the tautomers do matter!
Time for some calculations (B3LYP+GD3BJ/Def2-TZVPP/SCRF=water, using integral=(acc2e=14,grid=superfinegrid). The FAIR DOI for the collection is 10.14469/hpc/7990
In the diagram above, the non-aromatic valence representation is shown at the top, followed by a reorganisation of the electrons into an aromatic form below. These aromatic forms are all ionic with charge separation and so the question here arises: does the aromatic stabilisation energy outweigh the destabilisation caused by charge separation?
The calculated free energies (ΔG298) show that the oxime is 3.2 kcal/mol more stable than the next most stable, the hydroxylamine tautomer (calculated for a water continuum solvation model).The third tautomer is not far behind, being a nitrone! A 3D model is shown below illustrating the internal hydrogen bonds (I hope it is clear why I wanted the four stereocentres unambiguously identified, and why the simple perspective diagram shown on the Wikipedia page is not definitive). This model was obtained after also experimenting with hydroxyl rotamers to ensure the lowest energy was obtained.
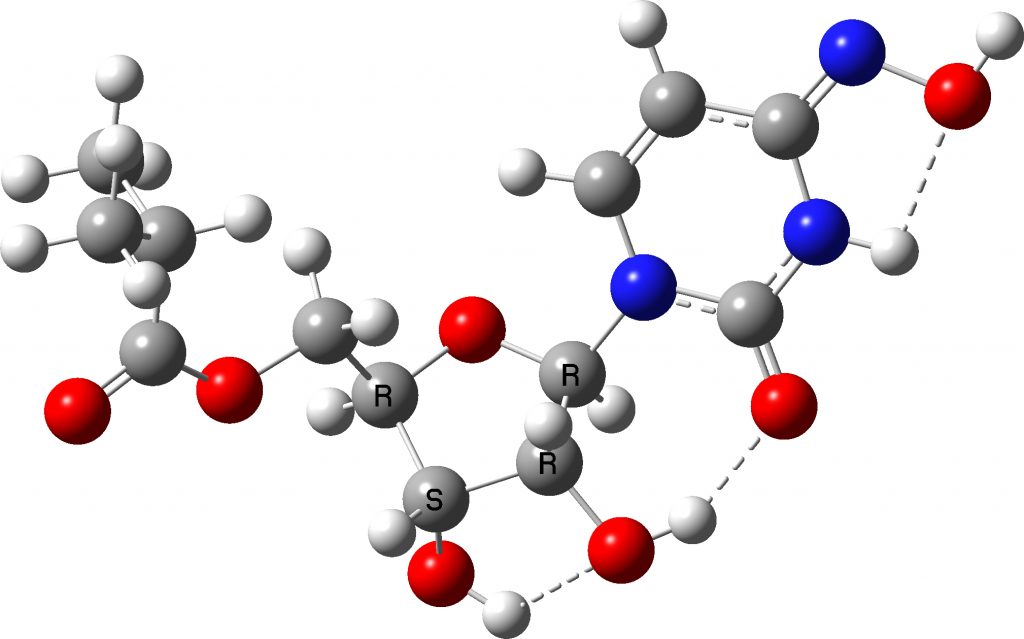
Click to load 3D model
So how “aromatic” are the two lowest energy species? The oxime has two sets of charge separation, whilst the hydroxylamine only one. One measure of aromaticity is the NICS magnetic index. For the oxime it is -0.1ppm, very clearly non-aromatic. Separating two sets of charges clearly is not compensated by aromatic stabilisation. The next most stable hydroxylamine has NICS -1.8ppm, which shows slightly more signs of aromaticity (benzene on this scale is ~ -10 ppm), perhaps because it only separates one pair of charges.
What about crystal structures? NETGEY is a model which removes the sugar unit and with an N-Me replacing the NH (thus preventing tautomerism and locking the molecule into the oxime form). MHCYTC protonates the hydroxylamine on the imine nitrogen OR the oxime on its imine nitrogen thus rendering the two tautomers identical.
| NETGEY | MHCYTC |
|---|---|
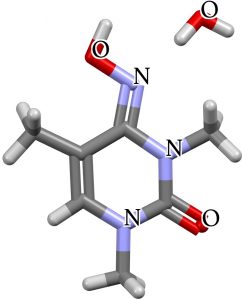 |
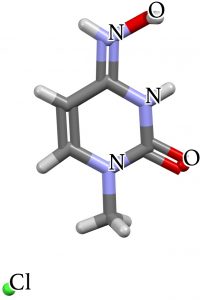 |
The crystal structure of Molnupiravir itself has not been reported, so there is no definitive answer to the most stable tautomer in the solid state. But the calculation above does suggest that it is the oxime (and hence SciFinder’s representation) that is the probable dominant form in aqueous solutions. If one is trying to build a model to show how this small molecule interacts with DNA, this might be useful information (in the same way that picking the correct tautomer of the original DNA bases worked for Watson and Crick!).
References
- J.D. WATSON, and F.H.C. CRICK, "Molecular Structure of Nucleic Acids: A Structure for Deoxyribose Nucleic Acid", Nature, vol. 171, pp. 737-738, 1953. https://doi.org/10.1038/171737a0
Today we have news that this molecule has been show to cut the risk of hospitalisation or death from Covid by about half; https://www.bbc.co.uk/news/health-58764440 and https://www.science.org/content/blog-post/molnupiravir-thor-s-hammer-delivers