We have been experimenting with full-colour 3D printing of molecular objects. I thought I might here share some of our observations. Firstly, I list the software used:
- Crystal structures as sources of ball&stick models (e.g. the CCDC database).
- Gaussian style cube files for sources of wavefunctions.
- There are plenty of other sources of course, in fact any file type which is supported by ..
- Jmol 13.3 (the app, not the applet) to render the above.
- And www.shapeways.com who provide the full colour 3D printing service.
My first attempt was to 3D print a molecular orbital; in fact the one shown in this post. To be printable, a 3D object must be fully connected, in other words not contain any disconnected components. To ensure this is true for a molecular orbital, one has to select a very low isosurface threshold; 0.005 au in this case. The orbital is computed as a Gaussian cube, and converted to a .wrl file as follows
- Start the Jmol.app.
- In the console, enter e.g.
isosurface sign color yellow green cutoff 0.005 "144.cub"
(where 144.cub is the Gaussian cube file). Then write model.wrl
to write out a VRML (.wrl) file.- Create an account at Shapeways, go to “make your own” and upload the .wrl file. After about five minutes, it should appear under “my models“, looking something like below:
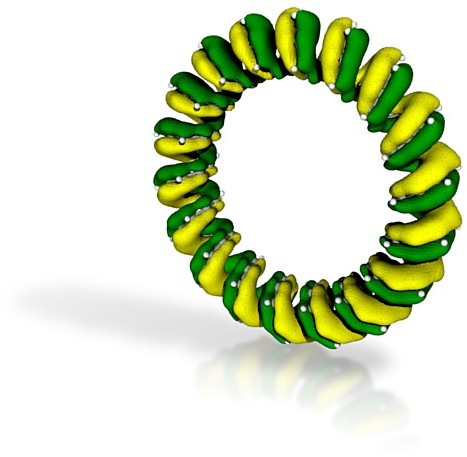

- Select “full color sandstone” as the material and see what the quoted price is. For the above it was €3.25. It is actually quite tricky to anticipate what physical size this represents, but upon printing and receiving it by post, it turned out to be about 2 cm in size! I should point out that it takes a day or so for Shapeways to confirm that the model has been sent to manufacturing (it is perfectly possible that it may be rejected at this stage because it cannot be manufactured for whatever reason).
- The thing now is to get the model to scale. To do this, a scale-factor has to be applied to the VRML file, which is easier said than done. Fortunately (thanks Bob Hanson!) this can be applied at the Jmol stage, and after step 2 above in the Jmol console, apply the command:
set exportScale 8.0
to scale up by 8-fold, before writing out the .wrl file. You know it has worked when the size is given as cm: 13.228 w x 13.288 d x 2.786 h and the price goes up to €69. I will assume (I have not printed at this scale yet) that it is less than 13 cm across, but something big enough to hand around a group of people. I should mention that we tried a variety of ways of scaling the model using programs such as Blender, but they all proved unreliable. You can of course also set the colours in the command above. - It is possible to set the scale larger than 8, but say at 10-fold the materials available to you reduce significantly. Full-colour sandstone is no longer available, and you can only print in a single colour (the materials by the way include stainless steel, for €5700.71).
- The resulting model is made from gypsum with a polymer binder. I have not had the courage yet to test how robust it is by breaking one of the models; I suspect it might be fairly fragile. Time will tell.
- I also tried a ball&stick. The molecule selected has been described in another post and is saved by converting the .cif file to .pdb (using in my case the program Mercury). This helps normalise the bond types, producing:
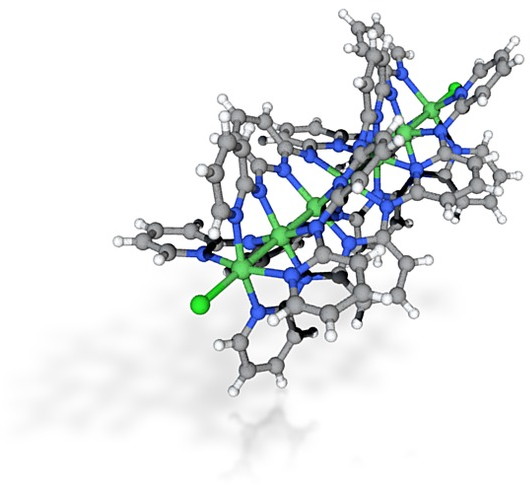
- In this case, converting the crystal coordinates via Jmol to .wrl required a scaling factor of 10.0 in this instance to produce a model costed as €24.53. It is very much a matter of adjusting the
exportscaleagainst the resulting printing price to get the desired result (by trial and error, but perhaps there is a more systematic way of doing this?). One of my colleagues (Paul) has had this sort of thing printed to about 25cm in size for about this price, and the model seems reasonably robust to physical handling. It may also benefit by fine-tuning of the bond radius; thicker would clearly be stronger.‡
If anyone reading this post has their own experience of 3D colour printing of chemical models, do please post comments here. And I dare say that in a few years time, students will simply press the “3D print” button on the tablet they are using to view lecture notes to get a copy. Mind you, I am somewhat ambivalent about such a process, having spent the last twenty years trying to discourage students from using the “2D print” button on their computer. Will I eventually come to adopt the same attitude to 3D print (if you click on the MO image above, you will get a virtual 3D model instead of a physical one).
‡ The width of a bond can be set using the Jmol command: set bondRadiusMilliAngstroms 300
Acknowledgments
This post has been cross-posted in PDF format at Authorea.
Tags: 3D colour printing, 3D printing, Bob Hanson, chemical models, EUR